Report with variants added by the user
The report can only include SNVs/Indels that have been added by the user. The method for adding variants to the report depends on the selected template type: "SNVs/Indels selected by user for reports" or "Pinned SNVs/Indels".
SNVs/Indels selected by user for reports#
The report is based on the report template block "SNVs/Indels selected by user for reports". In the block, you select the origin (germline or somatic) of variants that can be added to the report. If variants with the corresponding origin were discovered in the sample, the report will contain the message “You can mark variants to add them to this section in SNV Viewer”. How to include the required variants to the report is described here. How the included variants are presented in the report is described below. If the sample interpretation has been completed, but no variants have been included to the report, then the report will contain the message "Interpretation completed, no variants were added to the report". However, variants can be included to the report after the interpretation has been completed.
Remove variants from the report#
A variant can be removed from the report by hovering over its row and clicking
on . Then you need to
confirm removing in the opened window:
This variant will no longer be included to the report (activated
button ) on SNV Viewer page, too.
Pinned SNVs/Indels#
The report is based on the report template block "Pinned SNVs/Indels". In the block, you select the origin (germline or somatic) of variants that can be added to the report. If variants with the corresponding origin were discovered in the sample, the report will contain the message “You can pin variants in SNV Viewer to add them to this section”. To add a variant to the report, you need to pin it in the SNV Viewer table. To exclude a variant from the report, you need to unpin it on the SNV Viewer page. How the added variants are presented in the report is described below. If the sample interpretation has been completed, but no variants have been added to the report, then the report will contain the message "Interpretation completed, no variants were added to the report". However, variants can be added to the report after the interpretation has been completed.
How are the variants presented in the report?#
SNVs/Indels added to the report are presented in a table with the following columns:
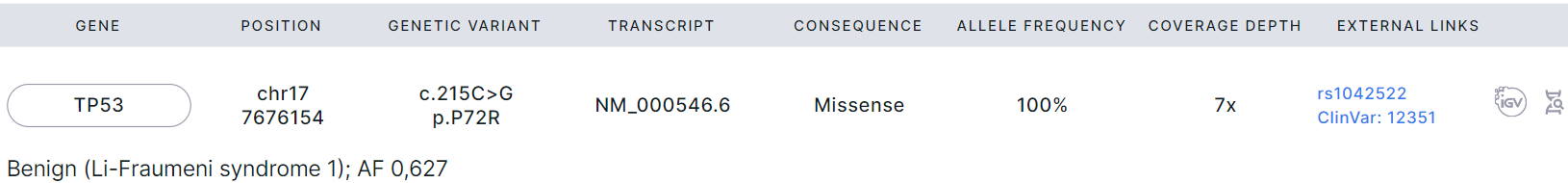
- Gene is the common name of the gene in which the variant is located. If you click on a gene, you will see a window with all the gene transcripts:

You can find the description of the transcripts' table columns in the description of "Transcripts" section on the variant details panel.
- Position is a coordinate of the variant in the genome (chromosome + start position).
- Genetic variant is the nucleotide and amino acid substitutions using the HGVS notation. Nucleotide substitution: “c.” (coding; for a substitution in the coding sequence) or “n.” (non-coding; for a substitution in the non-coding sequence) prefix + genomic position of the substituted nucleotide + reference allele > alternative allele. Amino acid substitution: “p.” prefix (protein) + reference amino acid + amino acid position in protein + new amino acid resulting from the substitution.
- Transcript is the main transcript ID from the RefSeq database (NM_xxxxxx.x). If you click on a transcript, you will see the same window with all the gene transcripts, as when you click on the "Gene" field.
- Consequence is the effect of the variant on genes. A detailed description of the possible values can be found here.
- Allele frequency is an alternative allele frequency for the sample (in percentages).
- Coverage depth is a sequencing depth; the total number of reads of the sequence overlapping the variant position for the sample.
- External links are links to pages with variant information in dbSNP, ClinVar and COSMIC (if it was uploaded as a custom annotation).
- Links to the variant in embedded services:
is a module for visualization of variant on the genome,
is a variant details page in SNV Viewer ("Annotation" tab).
The variant interpretation text may be given below the variant row if it was added as described here.
Reports export#
Reports with variants added by the user can be downloaded in PDF format.
To do this, click on the button in the
upper right corner of the report page.