Sample Interpretation
The sample interpretation is interpretation of the pathogenicity of mutations discovered in the sample. It is presented as interpretation results, i.e. information on the number of SNVs/Indels and/or copy number variations with a particular pathogenicity class discovered in the sample and included to the interpretation results.
Attention!
Do not confuse the sample interpretation described in this article with the variant interpretation or the variation interpretation. A variant or variation interpretation is a comment with important information about the mutation added by the user.
Where can I find the interpretation results?#
The sample interpretation results are displayed:
- on the "Samples" page (the "Results" column);
- on the sample page (the "Interpretation Results" section) - appear only after the interpretation is completed;
- on the "Sample" tab of the variant details page (the "Sample Info" section, the "Interpretation results" field);
- on the run page in the "Interpretation" column (for samples in a run).
What do the interpretation results look like?#
The sample interpretation results may include information about SNVs/Indels, designated "SNV", and information about copy number variations, designated "CNV". The information is presented as follows:
- pathogenicity icons indicating the number of mutations with this pathogenicity:
- pathogenic mutations:
;
- likely pathogenic mutations:
;
- mutations of uncertain significance:
;
- likely benign mutations:
;
- benign mutations:
.
- pathogenic mutations:
- message "Interpretation not completed" if mutations with a particular pathogenicity class were included to the interpretation results, but the sample interpretation was not completed.
- message "No pathogenic variants found" if the sample interpretation has been completed, but no mutations with a particular pathogenicity class have been included to the interpretation results.
How to complete and resume sample interpretation?#
You can complete and resume sample interpretation:
- on the "Samples" page;
- on the "Sample" tab of the variant details page;
- in the run SNV Viewer (for samples in a run).
Confirmation window#
During the sample interpretation completion by using any of the methods described below, you will see a confirmation window. For non-tumor samples, it looks like this:
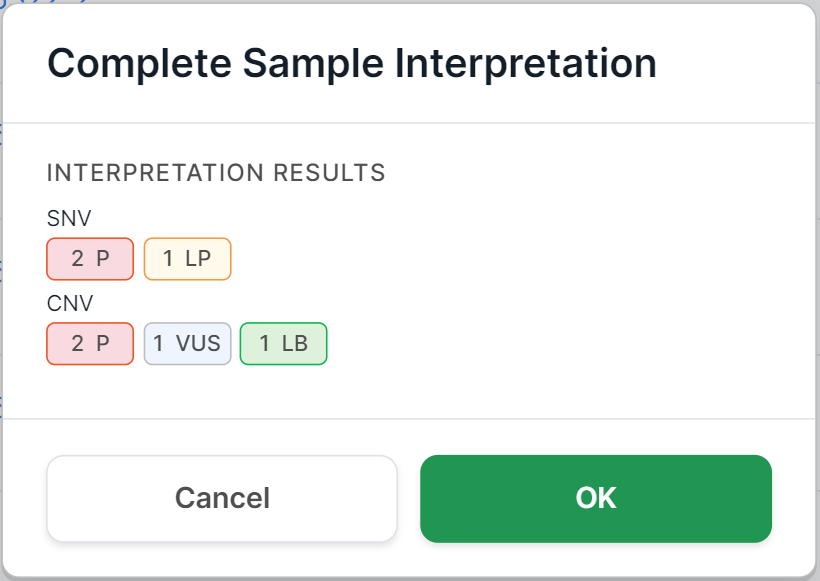
And for tumor samples, it also includes fields about the oncological disease:
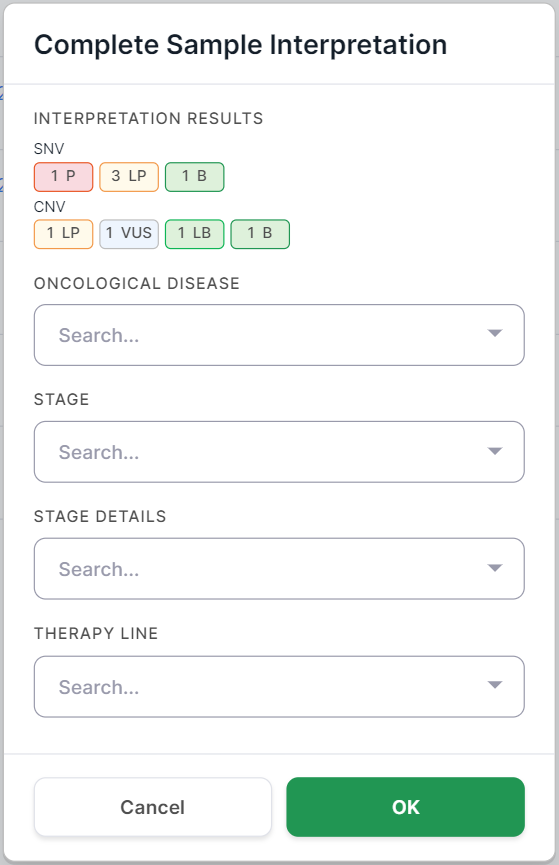
The information in the "Interpretation results" field will appear in the results once the interpretation is complete.
For tumor samples, you can specify in the confirmation window the oncological disease, its stage, and stage details which were defined for the patient, as well as the therapy line used to treat the patient's disease, at the time the sample was taken for sequencing.
I. Complete and resume interpretation on the "Samples" page#
This is the easiest way. First you need to open the "Samples" page (Genomenal home page).
Completion: Hover over the row of the sample whose interpretation you want to complete and click
on . Then click on
in the confirmation window.
Resuming: Hover over the sample row and
click on .
II. Complete and resume interpretation on the "Sample" tab#
To use this method, first you need to open the variant details page. If your goal is only to complete the sample
interpretation, a page of any variant discovered in that sample will do. This page can be opened from the sample
SNV Viewer page (to do this, click on the
button in any variant's row in the SNV Viewer table), or from the report page with SNVs/Indels (to do this, click on the button
in any variant's row in the report table).
On the page that opens, go to the "Sample" tab. There, in the “Sample Info” section, you will find the button that
you need to complete or resume the sample interpretation.
Completion: click on  . Then click on
. Then click on in the confirmation window.
Resuming: click on  .
.
III. Complete and resume interpretation in the run SNV Viewer#
This method is only available for samples included in the run. Open the run SNV Viewer and look for any variant discovered in the sample whose interpretation you want to complete (the sample name is listed in the "Sample" column).
Completion: click on the sample name in any variant's row and select the corresponding option:
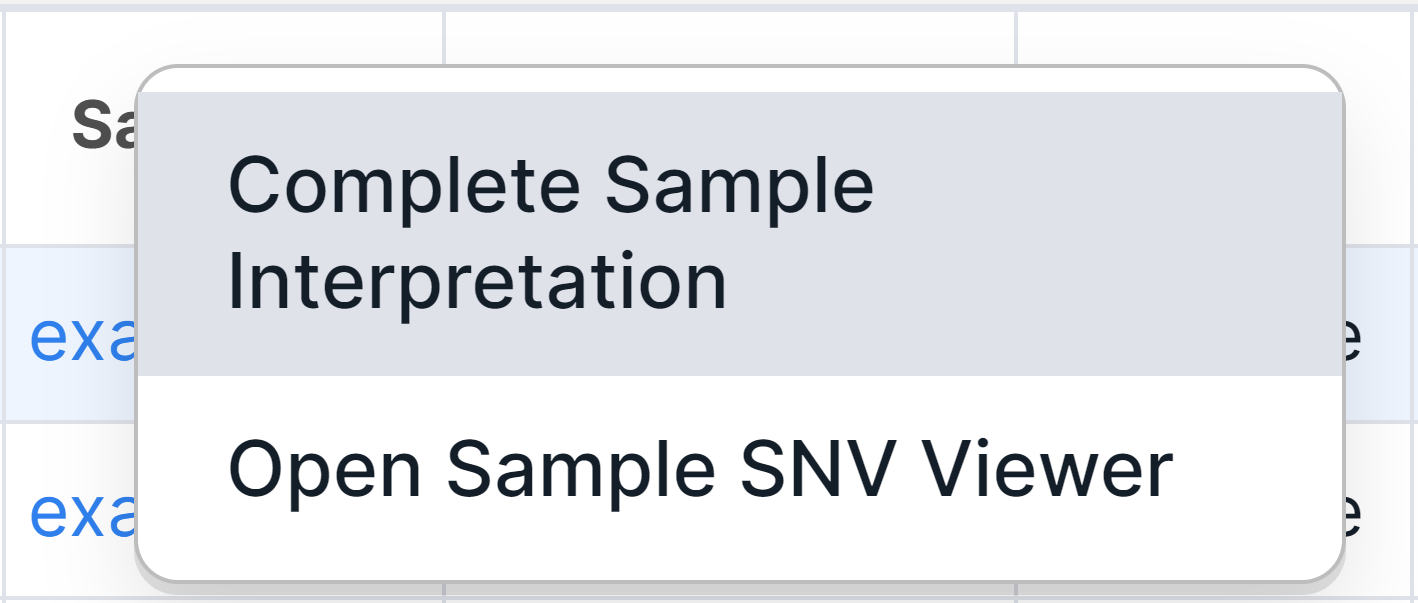
Then
click on in the confirmation window.
Run's samples with completed interpretation are
marked with :
Resuming: click on the sample name in any variant's row and select the corresponding option:

How to include a mutation to the interpretation results?#
To include SNV/Indel or copy number variation to the sample interpretation results, you need to:
- determine the mutation pathogenicity class (how to do this is described here for variants, and there for variations);
- include the mutation to interpretation results. This occurs automatically when the pathogenicity class of the mutation is determined. In addition, potential findings can be included to interpretation results, as described here.
note
If necessary, mutations can be added to the sample interpretation results after the interpretation is complete.