Structural variations discovery
After the "Variant discovery preprocessing" stage has been successfully completed, structural variations (SVs) discovery can be initiated for the samples if the stage is included in the analysis workflow. Germline structural variations will be discovered for a non-tumor sample and a single tumor sample, and somatic structural variations will be discovered for a tumor sample from the tumor/control sample set.
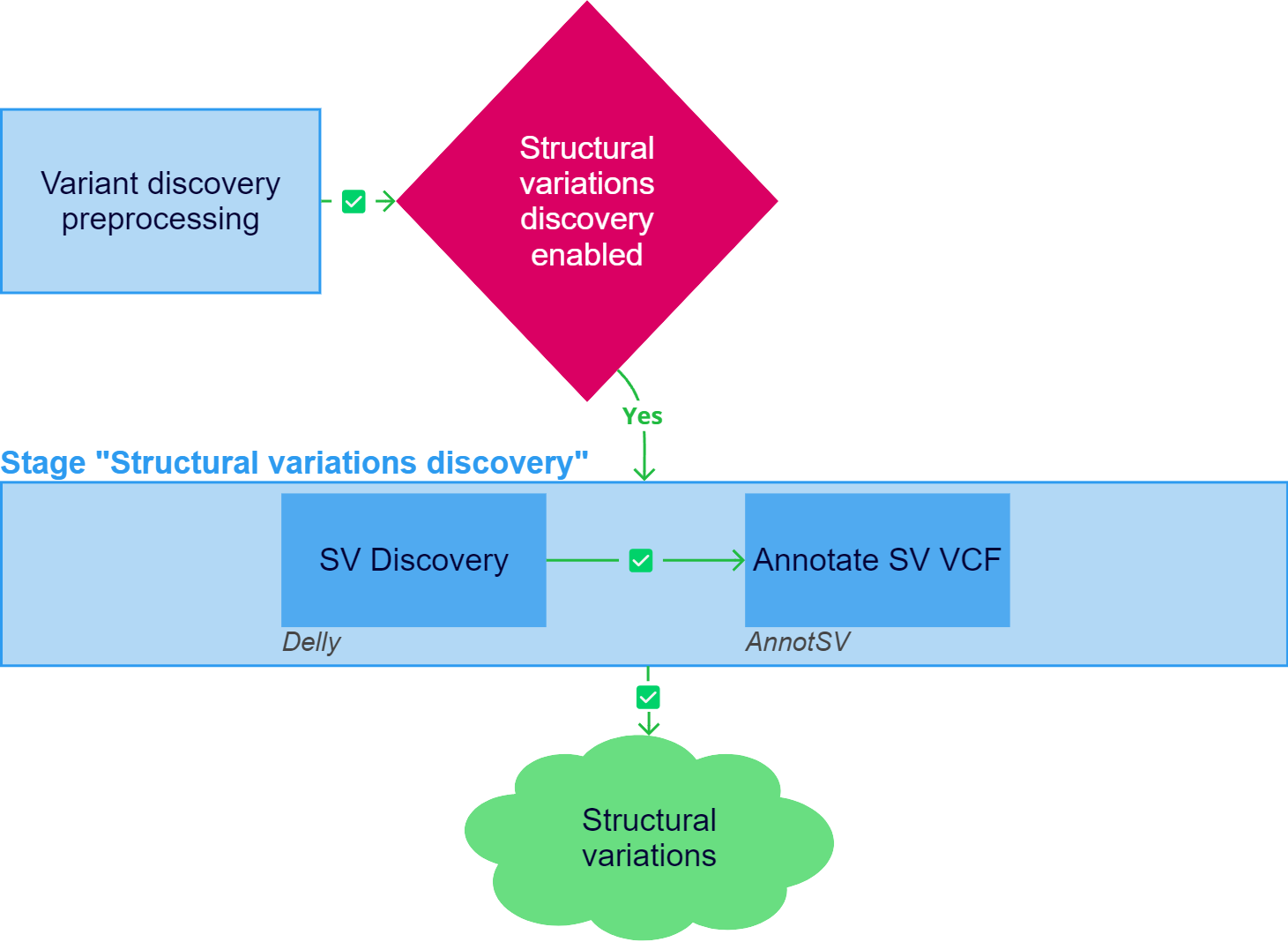
1. SV Discovery#
Structural variations calling is performed using the Delly tool. Delly is an integrated structural variation prediction method that can discover, genotype and visualize deletions, tandem duplications, inversions and translocations at single-nucleotide resolution in short-read and long-read massively parallel sequencing data. When calling structural variations, the telomere and centromere regions of chromosomes are not considered, since these repetitive regions cannot be analyzed using short-read data, as well as unplaced contigs.
To call somatic structural variations, at least one tumor sample and a matching normal sample are required. Therefore, somatic structural variations discovery is performed only in a tumor sample from a tumor/control sample set. In a single tumor sample, only germline structural variations are discovered.
Using the sample genome alignment (or the alignment of the tumor and non-tumor sample genomes in case of somatic structural variations discovery in the tumor sample from the tumor/control sample set), Delly calculates structural variations and outputs them as a BCF (binary variant call format) file, i.e. a binary version of VCF. The resulting file can be downloaded in the "Result files" section in the "SV Discovery" task details ("Download BCF"). The index file corresponding to this file can be downloaded in the same section ("Download CSI").
View, subset and filter VCF or BCF files by position and filtering expression. Convert between VCF and BCF. Former bcftools subset.
The resulting BCF file is then converted to a VCF file using bcftools view. The VCF file is then compressed into a GZIP archive using bgzip. The resulting file can be downloaded in the "Result files" section in the "SV Discovery" task details ("Download VCF_GZ"). You can also open this file in the IGV browser by clicking on the "Open in IGV Browser" link. In addition, the VCF file is indexed using tabix. The resulting index file can be downloaded in the same section ("Download VCF_TBI").
2. Annotate SV VCF#
Annotation and ranking of discovered structural variations is performed using the AnnotSV program. This tool compiles functionally, regulatory and clinically relevant information and aims at providing annotations useful to interpret SV potential pathogenicity and filter out SV potential false positives.
- The resulting file with annotated structural variations in TSV format can be downloaded in the "Result files" section in the "Annotate SV VCF" task details ("Download Filtered variants TSV"). You can also open it in Google Sheets. The same file in CSV format can be downloaded from the same section ("Download Filtered variants CSV").
- The resulting file with annotated structural variations in VCF format is compressed into a GZIP archive using bgzip. It can be downloaded in the same section ("Download Filtered variants VCF_GZ"). You can also open this file in the IGV browser by clicking on the "Open in IGV Browser" link. In addition, the VCF file is indexed using tabix. The resulting index file can be downloaded in the same section ("Download Filtered variants VCF_TBI").
info
If you want to add a track with structural variations discovered by the analysis of the sample you uploaded to Genomenal to your desktop IGV, you can do so via a link. Open the details of the necessary task ("SV Discovery" or "Annotate SV VCF") and do the following:
- Right-click the variation file link (depending on the selected task, this may be the link "Download VCF_GZ" or "Download Filtered variants VCF_GZ") and select the "Copy link address" option.
- Upload the track via URL to your desktop IGV, as described here.
- Right-click on the download link for the index file corresponding to the annotation file ("Download VCF_TBI" or "Download Filtered variants VCF_TBI") and select the "Copy link address" option.
- Add the index via URL to the corresponding field in IGV.
- Click "OK". Done! The track with structural variations discovered in the sample is added to IGV.
After the "Structural variations discovery" stage, the analysis can proceed with report generation.