Analysis of FASTQ or BAM samples
Files in FASTQ or BAM format can be uploaded as tumor samples, normal samples, or a set consisting of tumor sample(s) and a control sample.
The analysis of a sample set depends on its settings. They can be either set during composing the sample set as workflow settings, or changed on the "Parameters" tab after starting the analysis.
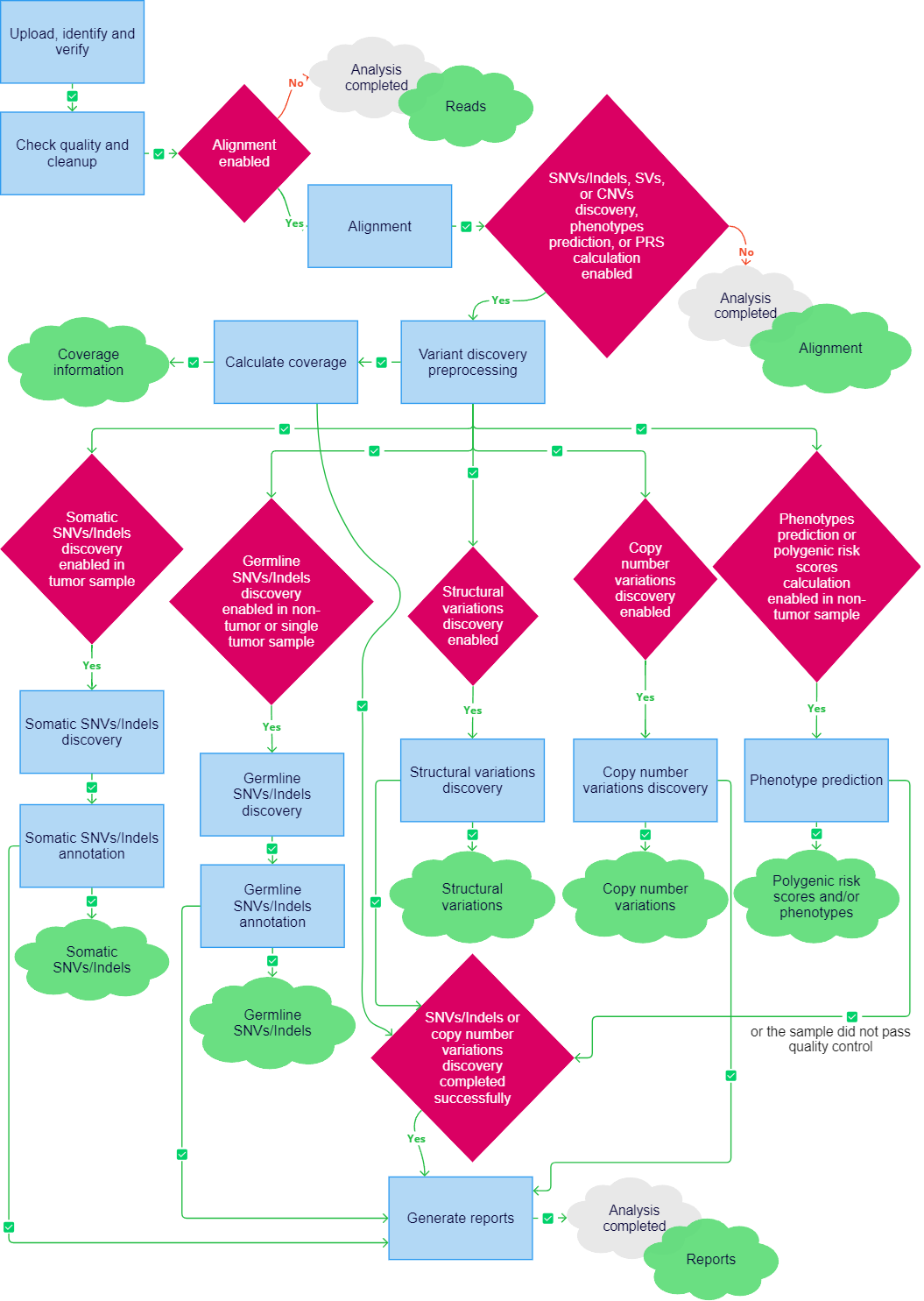
Analysis of a sample uploaded in FASTQ or BAM format may include the following stages:
- Upload, identify and verify;
- Check quality and cleanup;
- Alignment if alignment is included in the workflow;
- Variant discovery preprocessing if at least one of the following downstream stages is included in the workflow: SNVs/Indels discovery, structural variations discovery, copy number variations discovery, or at least one of the genomic predictions tasks (oligogenic/polygenic risk scores calculation, pharmacogenetic calculation, or ancestry analysis);
- Calculate coverage if at least one of the following downstream stages is included in the workflow: SNVs/Indels discovery, structural variations discovery, copy number variations discovery, or at least one of the genomic predictions tasks (oligogenic/polygenic risk scores calculation, pharmacogenetic calculation, or ancestry analysis);
- Somatic SNVs/Indels discovery if somatic SNVs/Indels discovery is included in the workflow and if sequencing sample is uploaded as tumor;
- Somatic SNVs/Indels annotation if somatic SNVs/Indels discovery is included in the workflow and if sequencing sample is uploaded as tumor;
- Germline SNVs/Indels discovery if germline SNVs/Indels discovery is included in the workflow and if sequencing sample is uploaded as normal or single tumor;
- Germline SNVs/Indels annotation if germline SNVs/Indels discovery is included in the workflow and if sequencing sample is uploaded as normal or single tumor;
- Structural variations discovery if structural variations discovery is included in the workflow;
- Copy number variations discovery if copy number variations discovery is included in the workflow and if a sequencing type and capture kit (if the sequencing type is defined as "WES" or "Panel") are detected for the sample;
- Genomic predictions if oligogenic risk scores calculation, polygenic risk scores calculation, pharmacogenetic calculation and/or ancestry analysis are included in the workflow;
- Generate reports if:
- At least one active report template, applicable to the sample type, with at least one added block was added to the system. The template can be applicable to tumor, non-tumor, or any sample type. A report template that meets all of the above conditions must be added to the system before the sample was processed. If you added or updated a report template later and want to generate or update the corresponding report for the sample, reprocess the sample analysis from the "Generate reports" stage.
- The sample workflow includes alignment and one of the following stages: "Somatic SNVs/Indels discovery" for a tumor sample, "Germline SNVs/Indels discovery" for a non-tumor or single tumor sample, or "Copy number variations discovery". If other stages are included in the workflow, it may only affect the generation of specific reports or their content.
- The sample analysis has been successfully completed (i.e. all stages included in the workflow have the status "Complete").